Welcome to easyEWAS!
Author: Yuting Wang, Xu Gao (Corresponding author)
Epigenome-Wide Association Study in R
easyEWAS is an R package for conducting EWAS in a unified and reproducible way. It supports Illumina methylation array platforms including 27K, 450K, EPIC v1, EPIC v2, and MSA, and provides an end-to-end workflow for modeling, batch correction, visualization, validation, enrichment, and optional DMR analysis.
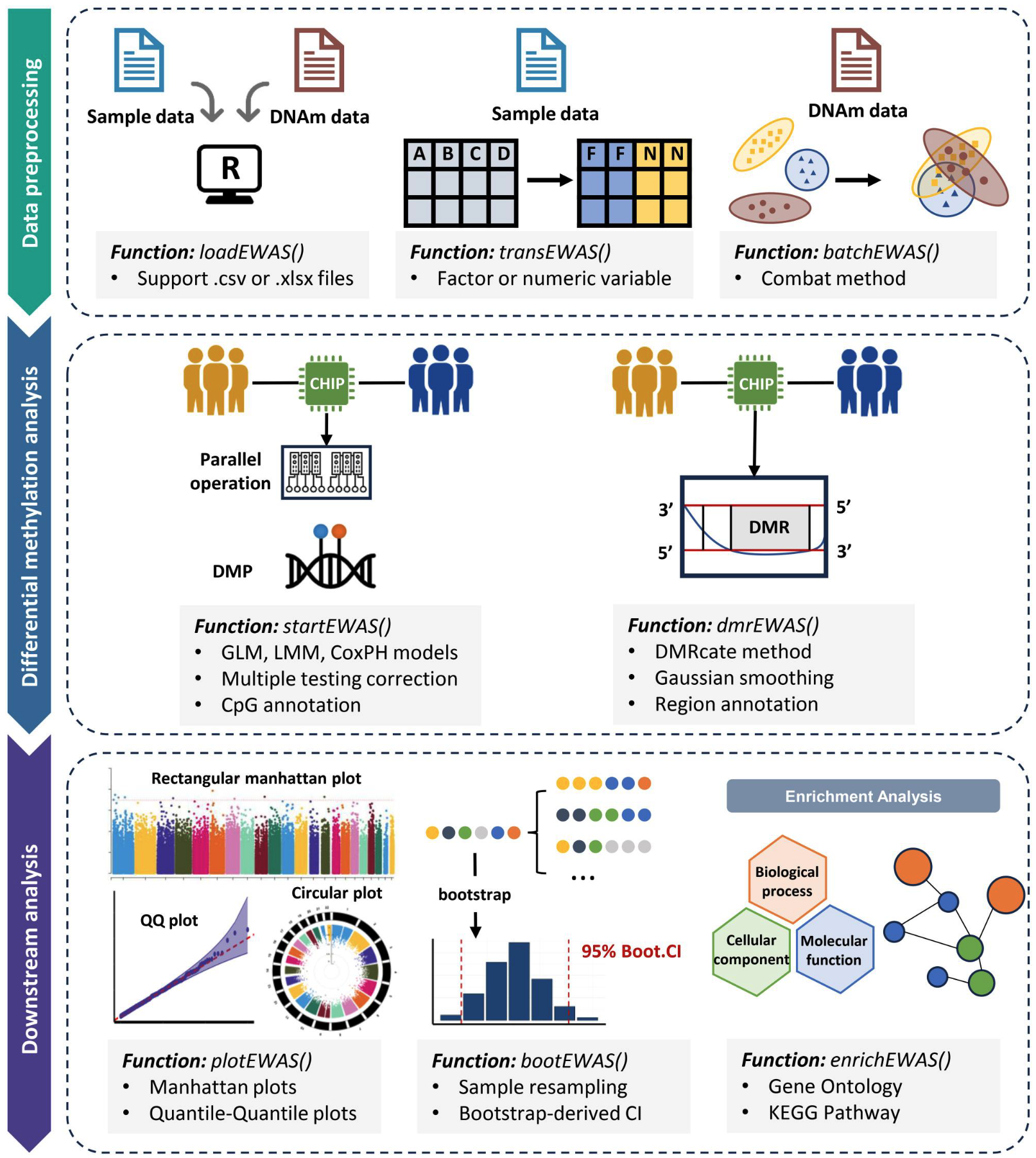
The Workflow Overview of easyEWAS
Installation
easyEWAS can now be installed from CRAN or GitHub. For
most users, we recommend the CRAN release. We also recommend
pre-installing optional dependencies used by advanced modules.
Install from CRAN (recommended)
install.packages("easyEWAS")Core dependencies by function
| Function | Required packages | Notes |
|---|---|---|
batchEWAS() |
sva |
Required for ComBat batch correction |
batchEWAS(..., parallel = TRUE) |
sva,
BiocParallel |
BiocParallel is needed only
for parallel mode |
enrichEWAS() |
clusterProfiler,
org.Hs.eg.db |
Required for ID conversion and GO/KEGG enrichment |
enrichEWAS(plot = TRUE, plotType = "dot") |
enrichplot |
Required for dotplot visualization |
dmrEWAS() |
DMRcate |
Required for DMR analysis |
Recommended pre-install
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
BiocManager::install(
c(
"sva",
"BiocParallel",
"clusterProfiler",
"org.Hs.eg.db",
"enrichplot",
"DMRcate"
),
ask = FALSE,
update = TRUE
)Install from GitHub (development version)
if (!requireNamespace("remotes", quietly = TRUE)) {
install.packages("remotes")
}
remotes::install_github("ytwangZero/easyEWAS")Load package
library(easyEWAS)If you prefer installing only when needed:
batchEWAS():BiocManager::install(c("sva", "BiocParallel"))enrichEWAS():BiocManager::install(c("clusterProfiler", "org.Hs.eg.db", "enrichplot"))dmrEWAS():BiocManager::install("DMRcate")
Citation
Wang Y, Jiang M, Niu S, Gao X. easyEWAS: a flexible and user-friendly R package for epigenome-wide association study. Bioinformatics Advances, 2025, 5(1): vbaf026.



